
ImmuHub® Platform Slected Publications
艾沐蒽ImmuHub®已累积发表
39篇
SCI论文
586.9
总影响因子

Measurable residual disease testing by next generation sequencing is more accurate compared with multiparameter flow cytometry in adults with B-cell acute lymphoblastic leukemia
Results of measurable residual disease (MRD)-testing by next-generation sequencing (NGS) correlate with relapse risk in adults with B-cell acute lymphoblastic leukemia (ALL) receiving chemotherapy or an allotransplant from a human leukocyte antigen (HLA)-identical relative or HLA-matched unrelated donor. We studied cumulative incidence of relapse (CIR) and survival prediction accuracy using a NGS-based MRD-assay targeting immunoglobulin genes after 2 courses of consolidation chemotherapy cycles in 93 adults with B-cell ALL most receiving HLA-haplotype-matched related transplants. Prediction accuracy was compared with MRD-testing using multi-parameter flow cytometry (MPFC). NGS-based MRD-testing detected residual leukemia in 28 of 65 subjects with a negative MPFC-based MRD-test. In Cox regression multi-variable analyses subjects with a positive NGS-based MRD-test had a higher 3-year CIR (Hazard Ratio [HR] = 3.37; 95 % Confidence Interval [CI], 1.34–8.5; P = 0.01) and worse survival (HR = 4.87 [1.53–15.53]; P = 0.007). Some data suggest a lower CIR and better survival in NGS-MRD-test-positive transplant recipients but allocation to transplant was not random. Our data indicate MRD-testing by NGS is more accurate compared with testing by MPFC in adults with B-cell ALL in predicting CIR and survival. (Registered in the Beijing Municipal Health Bureau Registration N 2007-1007 and in the Chinese Clinical Trial Registry [ChiCTR–OCH–10000940 and ChiCTROPC-14005546]).

Landscape of IGH germline genes of Chiroptera and the pattern of Rhinolophus affinis bat IGH CDR3 repertoire
The emergence and re-emergence of abundant viruses from bats that impact human and animal health have resulted in a resurgence of interest in bat immunology. Characterizing the immune receptor repertoire is critical to understanding how bats coexist with viruses in the absence of disease and developing new therapeutics to target viruses in humans and susceptible livestock. In this study, IGH germline genes of Chiroptera including Rhinolophus ferrumequinum, Phyllostomus discolor, and Pipistrellus pipistrellus were annotated, and we profiled the characteristics of Rhinolophus affinis (RA) IGH CDR3 repertoire. The germline genes of Chiroptera are quite different from those of human, mouse, cow, and dog in evolution, but the three bat species have high homology. The CDR3 repertoire of RA is unique in many aspects including CDR3 subclass, V/J genes access and pairing, CDR3 clones, and somatic high-frequency mutation compared with that of human and mouse, which is an important point in understanding the asymptomatic nature of viral infection in bats. This study unveiled a detailed map of bat IGH germline genes on chromosome level and provided the first immune receptor repertoire of bat, which will stimulate new avenues of research that are directly relevant to human health and disease.

Comprehensive application of AI algorithms with TCR NGS data for glioma diagnosis
T-cell receptor (TCR) detection can examine the extent of T-cell immune responses. Therefore, the article analyzed characteristic data of glioma obtained by DNA-based TCR high-throughput sequencing, to predict the disease with fewer biomarkers and higher accuracy. We downloaded data online and obtained six TCR-related diversity indices to establish a multidimensional classification system. By comparing actual presence of the 602 correlated sequences, we obtained two-dimensional and multidimensional datasets. Multiple classification methods were utilized for both datasets with the classification accuracy of multidimensional data slightly less to two-dimensional datasets. This study reduced the TCR β sequences through feature selection methods like RFECV (Recursive Feature Elimination with Cross-Validation). Consequently, using only the presence of these three sequences, the classification AUC value of 96.67% can be achieved. The combination of the three correlated TCR clones obtained at a source data threshold of 0.1 is: CASSLGGNTEAFF_TRBV12_TRBJ1-1, CASSYSDTGELFF_TRBV6_TRBJ2-2, and CASSLTGNTEAFF_TRBV12_TRBJ1-1. At 0.001, the combination is: CASSLGETQYF_TRBV12_TRBJ2-5, CASSLGGNQPQHF_TRBV12_TRBJ1-5, and CASSLSGNTIYF_TRBV12_TRBJ1-3. This method can serve as a potential diagnostic and therapeutic tool, facilitating diagnosis and treatment of glioma and other cancers.

New insights into the germline genes and CDR3 repertoire of the TCRβ chain in Chiroptera
Introduction: Bats are recognized as natural reservoirs for many viruses, and their unique immune system enables them to coexist with these viruses without frequently exhibiting disease symptoms. However, the current understanding of the bat adaptive immune system is limited due to the lack of a database or tool capable of processing T-cell receptor (TCR) sequences for bats. Methods: We performed germline gene annotation in three bat species using homologous genes and RSSs (Recombinational Signal Sequences) scanning method. Then we used the conserved C gene to construct the TCRβ chain receptor library of the Intermediate Horseshoe Bat. Bats' TCRβ data will be analyzed using MiXCR and constructed reference library. Results: Regarding the annotation results, we found that the Pale Spear-nosed Bat has 37 members in the TRBV12 family, which is more than the total number of TRBV genes in the Greater Horseshoe Bat. The average number of unique TCRβ chain receptor sequences in each Intermediate Horseshoe Bat sample reached 24,904. Discussion: The distinct variations in the distribution of TRBV genes among the three types of bats could have a direct impact on the diversity of the TCR repertoire, as evidenced by the presence of conserved amino acids that indicate the T-cell recognition of antigens in bats is MHC-restricted. The bats’ TCRβ repertoire is formed through the rearrangement of the V-D-J-C genes, with D-J/V-D deletions and insertions resulting in high diversity.

Increase in HIV reservoir and T cell immune response after CoronaVac vaccination in people living with HIV
【Introduction】 CoronaVac, an inactivated vaccine developed by Sinovac Life Sciences, has been widely used for protection against Coronavirus Disease 2019 (COVID-19). This study investigates its effect on the HIV reservoir and T cell repertoires in people living with HIV (PLWHs). 【Methods】 Blood samples were collected from fifteen PLWHs who were administered at least two doses of CoronaVac between April 2021 and February 2022. The levels of cell-associated HIV RNA (CA HIV RNA) and HIV DNA, as well as the T cell receptor (TCR) repertoire profiles, TCR clustering and TCRβ annotation, were studied. 【Results】 A significant increase was observed in CA HIV RNA at 2 weeks (431.5 ± 164.2 copies/106 cells, P = 0.039) and 12 weeks (330.2 ± 105.9 copies/106 cells, P = 0.019) after the second dose, when compared to the baseline (0 weeks) (73.6 ± 23.7 copies/106 cells). Various diversity indices of the TCRβ repertoire, including Shannon index, Pielou’s evenness index, and Hvj Index, revealed a slight increase (P<0.05) following CoronaVac vaccination. The proportion of overlapping TCRβ clonotypes increased from baseline (31.9%) to 2 weeks (32.5%) and 12 weeks (40.4%) after the second dose. We also found that the breadth and depth of COVID-19-specific T cells increased from baseline (0.003 and 0.0035) to 12 weeks (0.0066 and 0.0058) post the second dose. 【Conclusions】 Our study demonstrated an initial increase in HIV reservoir and TCR repertoire diversity, as well as an expansion in the depth and breadth of COVID-19-specific T-cell clones among CoronaVac-vaccinated PLWHs. These findings provide important insights into the effects of COVID-19 vaccination in PLWHs.

Sequential CD7 CAR T-Cell Therapy and Allogeneic HSCT without GVHD Prophylaxis
【BACKGROUND】 Patients with relapsed or refractory hematologic cancers have a poor prognosis. Chimeric antigen receptor (CAR) T-cell therapy as a bridge to allogeneic hematopoietic stem-cell transplantation (HSCT) has the potential for long-term tumor elimination. However, pre-HSCT myeloablation and graft-versus-host disease (GVHD) prophylaxis agents have toxic effects and could eradicate residual CAR T cells and compromise antitumor effects. Whether the integration of CAR T-cell therapy and allogeneic HSCT can preserve CAR T-cell function and improve tumor control is unclear. 【METHODS】 We tested a novel “all-in-one” strategy consisting of sequential CD7 CAR T-cell therapy and haploidentical HSCT in 10 patients with relapsed or refractory CD7-positive leukemia or lymphoma. After CAR T-cell therapy led to complete remission with incomplete hematologic recovery, patients received haploidentical HSCT without pharmacologic myeloablation or GVHD prophylaxis drugs. Toxic effects and efficacy were closely monitored. 【RESULTS】 After CAR T-cell therapy, all 10 patients had complete remission with incomplete hematologic recovery and grade 4 pancytopenia. After haploidentical HSCT, 1 patient died on day 13 of septic shock and encephalitis, 8 patients had full donor chimerism, and 1 patient had autologous hematopoiesis. Three patients had grade 2 HSCT-associated acute GVHD. The median follow-up was 15.1 months (range, 3.1 to 24.0) after CAR T-cell therapy. Six patients remained in minimal residual disease–negative complete remission, 2 had a relapse of CD7-negative leukemia, and 1 died of septic shock at 3.7 months. The estimated 1-year overall survival was 68% (95% confidence interval [CI], 43 to 100), and the estimated 1-year disease-free survival was 54% (95% CI, 29 to 100). 【CONCLUSIONS】 Our findings suggest that sequential CD7 CAR T-cell therapy and haploidentical HSCT is safe and effective, with remission and serious but reversible adverse events. This strategy offers a feasible approach for patients with CD7-positive tumors who are ineligible for conventional allogeneic HSCT. (Funded by the National Natural Science Foundation of China and the Key Project of Science and Technology Department of Zhejiang Province; ClinicalTrials.gov numbers, NCT04599556 and NCT04538599.)

Establishment of a humanized mouse model using steady-state peripheral blood-derived hematopoietic stem and progenitor cells facilitates screening of cancer-targeted T-cell repertoires
【Background】 Cancer-targeted T-cell receptor T (TCR-T) cells hold promise in treating cancers such as hematological malignancies and breast cancers. However, approaches to obtain cancer-reactive TCR-T cells have been unsuccessful. 【Methods】 Here, we developed a novel strategy to screen for cancer-targeted TCR-T cells using a special humanized mouse model with person-specific immune fingerprints. Rare steady-state circulating hematopoietic stem and progenitor cells were expanded via three-dimensional culture of steady-state peripheral blood mononuclear cells, and then the expanded cells were applied to establish humanized mice. The human immune system was evaluated according to the kinetics of dendritic cells, monocytes, T-cell subsets, and cytokines. To fully stimulate the immune response and to obtain B-cell precursor NAML-6- and triple-negative breast cancer MDA-MB-231-targeted TCR-T cells, we used the inactivated cells above to treat humanized mice twice a day every 7 days. Then, human T cells were processed for TCR β-chain (TRB) sequencing analysis. After the repertoires had been constructed, features such as the fraction, diversity, and immune signature were investigated. 【Results】 The results demonstrated an increase in diversity and clonality of T cells after treatment. The preferential usage and features of TRBV, TRBJ, and the V–J combination were also changed. The stress also induced highly clonal expansion. Tumor burden and survival analysis demonstrated that stress induction could significantly inhibit the growth of subsequently transfused live tumor cells and prolong the survival of the humanized mice. 【Conclusions】 We constructed a personalized humanized mouse model to screen cancer-targeted TCR-T pools. Our platform provides an effective source of cancer-targeted TCR-T cells and allows for the design of patient-specific engineered T cells. It therefore has the potential to greatly benefit cancer treatment.
An oncolytic virus delivering tumor-irrelevant bystander T cell epitopes induces anti-tumor immunity and potentiates cancer immunotherapy
Tumor-specific T cells are crucial in anti-tumor immunity and act as targets for cancer immunotherapies. However, these cells are numerically scarce and functionally exhausted in the tumor microenvironment (TME), leading to inefficacious immunotherapies in most patients with cancer. By contrast, emerging evidence suggested that tumor-irrelevant bystander T (TBYS) cells are abundant and preserve functional memory properties in the TME. To leverage TBYS cells in the TME to eliminate tumor cells, we engineered oncolytic virus (OV) encoding TBYS epitopes (OV-BYTE) to redirect the antigen specificity of tumor cells to pre-existing TBYS cells, leading to effective tumor inhibition in multiple preclinical models. Mechanistically, OV-BYTE induced epitope spreading of tumor antigens to elicit more diverse tumor-specific T cell responses. Remarkably, the OV-BYTE strategy targeting human severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)-specific T cell memory efficiently inhibited tumor progression in a human tumor cell-derived xenograft model, providing important insights into the improvement of cancer immunotherapies in a large population with a history of SARS-CoV-2 infection or coronavirus disease 2019 (COVID-19) vaccination.

Neutrophil activation and clonal CAR-T re-expansion underpinning cytokine release syndrome during ciltacabtagene autoleucel therapy in multiple myeloma
Cytokine release syndrome (CRS) is the most common complication of chimeric antigen receptor redirected T cells (CAR-T) therapy. CAR-T toxicity management has been greatly improved, but CRS remains a prime safety concern. Here we follow serum cytokine levels and circulating immune cell transcriptomes longitudinally in 26 relapsed/refractory multiple myeloma patients receiving the CAR-T product, ciltacabtagene autoleucel, to understand the immunological kinetics of CRS. We find that although T lymphocytes and monocytes/macrophages are the major overall cytokine source in manifest CRS, neutrophil activation peaks earlier, before the onset of severe symptoms. Intracellularly, signaling activation dominated by JAK/STAT pathway occurred prior to cytokine cascade and displayed regular kinetic changes. CRS severity is accurately described and potentially predicted by temporal cytokine secretion signatures. Notably, CAR-T re-expansion is found in three patients, including a fatal case characterized by somatic TET2-mutation, clonal expanded cytotoxic CAR-T, broadened cytokine profiles and irreversible hepatic toxicity. Together, our findings show that a latent phase with distinct immunological changes precedes manifest CRS, providing an optimal window and potential targets for CRS therapeutic intervention and that CAR-T re-expansion warrants close clinical attention and laboratory investigation to mitigate the lethal risk.
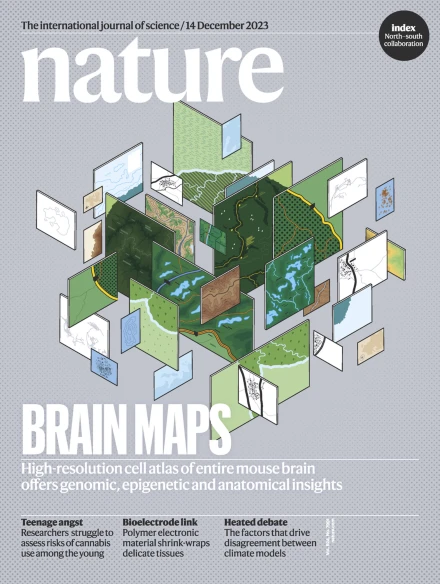

Inhaled SARS-CoV-2 vaccine for single-dose dry powder aerosol immunization
The COVID-19 pandemic has fostered major advances in vaccination technologies1,2,3,4; however, there are urgent needs for vaccines that induce mucosal immune responses and for single-dose, non-invasive administration4,5,6. Here we develop an inhalable, single-dose, dry powder aerosol SARS-CoV-2 vaccine that induces potent systemic and mucosal immune responses. The vaccine encapsulates assembled nanoparticles comprising proteinaceous cholera toxin B subunits displaying the SARS-CoV-2 RBD antigen within microcapsules of optimal aerodynamic size, and this unique nano–micro coupled structure supports efficient alveoli delivery, sustained antigen release and antigen-presenting cell uptake, which are favourable features for the induction of immune responses. Moreover, this vaccine induces strong production of IgG and IgA, as well as a local T cell response, collectively conferring effective protection against SARS-CoV-2 in mice, hamsters and nonhuman primates. Finally, we also demonstrate a mosaic iteration of the vaccine that co-displays ancestral and Omicron antigens, extending the breadth of antibody response against co-circulating strains and transmission of the Omicron variant. These findings support the use of this inhaled vaccine as a promising multivalent platform for fighting COVID-19 and other respiratory infectious diseases.
The effectiveness of blinatumomab in clearing measurable residual disease in pediatric B-cell acute lymphoblastic leukemia patients detected by next-generation sequencing
Background Blinatumomab improved survival outcomes in B-cell acute lymphoblastic leukemia (B-ALL) patients with measurable residual disease (MRD) <10−4. However, data on blinatumomab clearing MRD with high sensitivity of 10−6 remain scarce. This study evaluates the effectiveness of blinatumomab in eradicating extremely low level (up to <10−6) of MRD, as detected by next-generation sequencing (NGS), in children with B-ALL. Methods Patients (n = 19) whose MRD was undetectable by multiparameter flow cytometry (MFC) (sensitivity of 10−4) but detectable by NGS after chemotherapy and followed by blinatumomab consolidation were included retrospectively. Results After one course of blinatumomab, 13/19 patients (68%) successfully achieved NGS-MRD clearance (undetectable). With a median follow-up of 13.3 months, three of patients who were NGS-MRD positive relapsed within 1.8 months, while another three remained complete remission. Conclusions Our study was the first to demonstrate that blinatumomab could further eradicate MRD after patients achieve MFC-MRD undetectable in B-ALL patients.

Minimal residual disease detection by next-generation sequencing of different immunoglobulin gene rearrangements in pediatric B-ALL
While the prognostic role of immunoglobulin heavy chain locus (IGH) rearrangement in minimal residual disease (MRD) in pediatric B-acute lymphoblastic leukemia (B-ALL) has been reported, the contribution of light chain loci (IGK/IGL) remains elusive. This study is to evaluate the prognosis of IGH and IGK/IGL rearrangement-based MRD detected by next-generation sequencing in B-ALL at the end of induction (EOI) and end of consolidation (EOC). IGK/IGL rearrangements identify 5.5% of patients without trackable IGH clones. Concordance rates for IGH and IGK/IGL are 79.9% (cutoff 0.01%) at EOI and 81.0% (cutoff 0.0001%) at EOC, respectively. Patients with NGS-MRD < 0.01% at EOI or <0.0001% at EOC present excellent outcome, with 3-year event-free survival rates higher than 95%. IGH-MRD is prognostic at EOI/EOC, while IGK-MRD at EOI/EOC and IGL-MRD at EOI are not. At EOI, NGS identifies 26.2% of higher risk patients whose MRD < 0.01% by flow cytometry. However, analyzing IGK/IGL along with IGH fails to identify additional higher risk patients both at EOI and at EOC. In conclusion, IGH is crucial for MRD monitoring while IGK and IGL have relatively limited value.

T-cell receptor sequencing reveals hepatocellular carcinoma immune characteristics according to Barcelona Clinic liver cancer stages within liver tissue and peripheral blood
Analysis of T-cell receptor (TCR) repertoires in different stages of hepatocellular carcinoma (HCC) might help to elucidate its pathogenesis and progression. This study aimed to investigate TCR profiles in liver biopsies and peripheral blood mononuclear cells (PBMCs) in different Barcelona Clinic liver cancer (BCLC) stages of HCC. Ten patients in early stage (BCLC_A), 10 patients in middle stage (BCLC_B), and 10 patients in late stage (BCLC_C) cancer were prospectively enrolled. The liver tumor tissues, adjacent tissues, and PBMCs of each patient were collected and examined by TCR β sequencing. Based on the ImMunoGeneTics (IMGT) database, we aligned the V, D, J, and C gene segments and identified the frequency of CDR3 sequences and amino acids sequence. Diversity of TCR in PBMCs was higher than in both tumor tissues and adjacent tissues, regardless of BCLC stage and postoperative recurrence. TCR clonality was increased in T cells from peripheral blood in advanced HCC, compared with the early and middle stages. No statistical differences were observed between different BCLC stages, either in tumors or adjacent tissues. TCR clonality revealed no significant difference between recurrent tumor and non-recurrent tumor, therefore PBMCs was better to be representative of TCR characteristics in different stages of HCC compared to tumor tissues. Clonal expansion of T cells was associated with low risk of recurrence in HCC patients.

The Effectiveness of Blinatumomab in Clearing Next-Generation Sequencing Measurable Residual Disease in Pediatric Patients with B-Cell Acute Lymphoblastic Leukemia
Blinatumomab is a bispecific T-cell engager antibody designed to redirect CD3+ T cells to bind to CD19+ target cells in relapsed/refractory B-cell acute lymphoblastic leukemia (B-ALL). Measurable residual disease (MRD) is a significant prognostic factor for relapse and overall survival in B-ALL. Blinatumomab consolidation therapy has been shown to improve MRD remission (<10 -4), as detected by multiparameter flow cytometry (MFC) or reverse transcriptase-polymerase chain reaction (PCR), and survival outcomes in children with high-risk first relapsed B-ALL. The next-generation sequencing (NGS)-MRD is a highly sensitive technique that can detect very low levels of MRD (<10 -6) in patients who were considered “MRD negative” by less sensitive MFC or PCR technologies. Early achievement of NGS-MRD negativity (<10 -6) can help in identifying patients who have a very low risk of relapse. It has been suggested that blinatumomab treatment could help to attain deeper responses than that measured with a MRD sensitivity level of 10 -4. However, data on blinatumomab clearing MRD with high sensitivity of 10 -6 remain scarce. Hence, we evaluated the effectiveness of blinatumomab in eradicating extremely low level of MRD (<10 -6), as detected by NGS, in children with B-ALL.

Generation of whole tumor cell vaccine for on-demand manipulation of immune responses against cancer under near-infrared laser irradiation
The therapeutic efficacy of whole tumor cell vaccines (TCVs) is modest, which has delayed their translation into personalized immunotherapies in the clinic. Here, we develop a TCV platform based on photothermal nanoparticle-loaded tumor cells, which can be rationally applied to diverse tumor types to achieve on-demand boost of anti-tumor immune responses for inhibiting tumor growth. During the fabrication process, mild photothermal heating by near-infrared (NIR) laser irradiation induces the nanoparticle-bearing tumor cells to express heat shock proteins as endogenous adjuvants. After a single vaccination at the back of tumor-bearing mice, non-invasive NIR laser irradiation further induces mild hyperthermia at vaccination site, which promotes the recruitment, activation, and antigen presentation by dendritic cells. Using an indicator we term fluctuation of tumor growth rate, we determine appropriate irradiation regimens (including optimized irradiation intervals and times). This TCV platform enables on-demand NIR manipulation of immune responses, and we demonstrate potent therapeutic efficacy against six murine models that mimick a range of clinical scenarios, including a model based on humanized mice and patient-derived tumor xenografts.

Applying T-classifier, binary classifiers, upon high-throughput TCR sequencing output to identify cytomegalovirus exposure history
With the continuous development of information technology and the running speed of computers, the development of informatization has led to the generation of increasingly more medical data. Solving unmet needs such as employing the constantly developing artificial intelligence technology to medical data and providing support for the medical industry is a hot research topic. Cytomegalovirus (CMV) is a kind of virus that exists widely in nature with strict species specificity, and the infection rate among Chinese adults is more than 95%. Therefore, the detection of CMV is of great importance since the vast majority of infected patients are in a state of invisible infection after the infection, except for a few patients with clinical symptoms. In this study, we present a new method to detect CMV infection status by analyzing high-throughput sequencing results of T cell receptor beta chains (TCRβ). Based on the high-throughput sequencing data of 640 subjects from cohort 1, Fisher’s exact test was performed to evaluate the relationship between TCRβ sequences and CMV status. Furthermore, the number of subjects with these correlated sequences to different degrees in cohort 1 and cohort 2 were measured to build binary classifier models to identify whether the subject was CMV positive or negative. We select four binary classification algorithms: logistic regression (LR), support vector machine (SVM), random forest (RF), and linear discriminant analysis (LDA) for side-by-side comparison. According to the performance of different algorithms corresponding to different thresholds, four optimal binary classification algorithm models are obtained. The logistic regression algorithm performs best when Fisher's exact test threshold is 10−5, and the sensitivity and specificity are 87.5% and 96.88%, respectively. The RF algorithm performs better at the threshold of 10−5, with a sensitivity of 87.5% and a specificity of 90.63%. The SVM algorithm also achieves high accuracy at the threshold value of 10−5, with a sensitivity of 85.42% and specificity of 96.88%. The LDA algorithm achieves high accuracy with 95.83% sensitivity and 90.63% specificity when the threshold value is 10−4. This is probably because the two-dimensional distribution of CMV data samples is linearly separable, and linear division models such as LDA are more effective, while the division effect of nonlinear separable algorithms such as random forest is relatively inaccurate. This new finding may be a potential diagnostic method for CMV and may even be applicable to other viruses, such as the infectious history detection of the new coronavirus.

New insights into the germline genes and CDR3 repertoire of the TCRβ chain in Chiroptera
Introduction: Bats are recognized as natural reservoirs for many viruses, and their unique immune system enables them to coexist with these viruses without frequently exhibiting disease symptoms. However, the current understanding of the bat adaptive immune system is limited due to the lack of a database or tool capable of processing T-cell receptor (TCR) sequences for bats. Methods: We performed germline gene annotation in three bat species using homologous genes and RSSs (Recombinational Signal Sequences) scanning method. Then we used the conserved C gene to construct the TCRβ chain receptor library of the Intermediate Horseshoe Bat. Bats' TCRβ data will be analyzed using MiXCR and constructed reference library. Results: Regarding the annotation results, we found that the Pale Spear-nosed Bat has 37 members in the TRBV12 family, which is more than the total number of TRBV genes in the Greater Horseshoe Bat. The average number of unique TCRβ chain receptor sequences in each Intermediate Horseshoe Bat sample reached 24,904. Discussion: The distinct variations in the distribution of TRBV genes among the three types of bats could have a direct impact on the diversity of the TCR repertoire, as evidenced by the presence of conserved amino acids that indicate the T-cell recognition of antigens in bats is MHC-restricted. The bats’ TCRβ repertoire is formed through the rearrangement of the V-D-J-C genes, with D-J/V-D deletions and insertions resulting in high diversity.

Predictive value of next-generation sequencing-based minimal residual disease after CAR-T cell therapy
Minimal residual disease (MRD) is closely associated with risk stratification of hematological malignancies. Chimeric antigen receptor T (CAR-T) cell therapy, specifically redirects genetically modified immune cells to fight against hematologic malignancies. Studies have demonstrated that nearly 80–90% of patients with acute lymphoblastic leukemia (ALL) achieve complete remission (CR) after CAR-T cell therapy ; however, the 1-year relapse rate is still as high as 50–60%. Given the situation, nextgeneration sequencing (NGS)—MRD can accurately and quantitatively monitor patient-specific tumor cells (up to 10E−6) to reveal accurate and highly sensitive MRD levels. In this retrospective study, we estimated the sensitivity and concordance of NGS-MRD and FC-MRD of 20 patients with relapsed/refractory B-ALL who achieved CR after CAR-T cell therapy and further demonstrated the prognostic value of post-CART NGS-MRD at day 30 for ALL patients receiving CAR-T cell therapy.

Characteristics of T Cell Receptor Beta Repertoires Associated With Behçet’s Syndrome Patients With Major Organ Involvements
Background: T lymphocytes are one of the major components of the adaptive immunity in Behçet’s syndrome (BS) pathology. To further understand of the role of T cells in Behçet’s syndrome (BS), we explore disease related T cell receptor (TCR) repertoires. Methods: We performed a bulk sequencing of the TCR beta chain (TRB) of peripheral T-cells collected from 45 BS patients with panuveitis, intestinal involvement, and cardiovascular lesions and 10 cases of healthy controls (HC). Data analysis included peptide sequences, diversity analysis, variable (V)-joining (J) gene usage and K-nearest neighbor algorithm. Results: We found significant differences in V, J and V-J combinations between the BS patients and HC. The decrease in TCR clone diversity indicates clonal expansion in BS. Although no significant difference in TCR clone diversity was observed among patient subgroups, the patients with panuveitis displayed the highest heterogeneous TCR distribution. In addition, a set of V-J genes could effectively discriminate between active and inactive BS patients with an area under the receiver operating characteristics (ROC) curve of 0.88 (95% CI: 0.77-1.00). Conclusions: Clonal T cell expansion has been observed in BS patients. TCR profiles could help in the discrimination between active and inactive BS.

The Relationship Between Convergent IGH Signatures and Severity of COVID-19 Patients by Next-Generation Sequencing of B-Cell Repertoire
【Object】To reveal convergent IGH signatures and the association with severity of coronavirus disease 2019 (COVID-19) patients. 【Method】A total of 25 COVID-19 inpatients were classified into three clinical conditions: mild, severe, and critical. We analyzed convergent IGH signatures by ImmuHub® B-cell receptor (BCR) profiling system. 【Results】IGH singleton frequency in patients is significantly lower than that of healthy donors (HDs). The clonality index of IGH in patients is significantly higher than that in HDs. Nevertheless, no significant difference was observed among the three groups. The difference in IGH clonality (top five clones) between post- and pretreatment was significant in the improvement and deterioration groups. Three common public motifs were shared by all COVID-19 patients: ARDYGG, RWYFDY, and YYYYGMDV. 【Conclusion】 B cells could recognize severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and produce clonal expansion. Patients who had better outcomes after treatment had higher IGH clonality. Three common public motifs—ARDYGG, RWYFDY, and YYYYGMDV—might be used for vaccine development (ChiCTR2000029626).

Encephalomyelitis Caused by Balamuthia mandrillaris in a Woman With Breast Cancer: A Case Report and Review of the Literature
Balamuthia mandrillaris is one cause of a rare and severe brain infection called granulomatous amoebic encephalitis (GAE), which has a mortality rate of >90%. Diagnosis of Balamuthia GAE is difficult because symptoms are non-specific. Here, we report a case of Balamuthia amoebic encephalomyelitis (encephalitis and myelitis) in a woman with breast cancer. She sustained trauma near a garbage dump 2 years ago and subsequently developed a skin lesion with a Mycobacterium abscessus infection. She experienced dizziness, lethargy, nausea and vomiting, inability to walk, and deterioration of consciousness. Next-generation sequencing of cerebrospinal fluid (CSF) samples revealed B. mandrillaris, and MRI of both brain and spinal cord showed abnormal signals. T-cell receptor (TCR) sequencing of the CSF identified the Top1 TCR. A combination of amphotericin B, flucytosine, fluconazole, sulfamethoxazole, trimethoprim, clarithromycin, pentamidine, and miltefosine was administrated, but she deteriorated gradually and died on day 27 post-admission.

Predictive Value of Next-Generation Sequencing-Based Minimal Residual Disease after CAR-T Cell Therapy
Minimal residual disease (MRD) is closely associated with risk stratification of hematological malignancies. Monitoring the MRD levels of patients during treatment and at time points after remission is critical for prevention of relapse. Chimeric antigen receptor T (CAR-T) cell therapy redirects genetically modified immune cells to fight against hematologic malignancies. However, given the high relapse rate after CAR-T cell therapy, MRD monitoring by traditional techniques cannot accurately quantify the disease burden, nor can they perform high-sensitivity in-depth monitoring. Further treatment options are still controversial at such a time point when the flow cytometry (FC) results show MRD < 0.01%, while tumor clones remain. Next-generation sequencing (NGS) - MRD screens out patient-specific T/B cell receptors and accurately and quantitatively monitor patient-specific tumor cells (up to 10E-6) to reveal accurate and highly sensitive MRD levels, plus provide more timely intervention criteria.
Specific CD8+ TCR Repertoire Recognizing Conserved Antigens of SARS-CoV-2 in Unexposed Population: A Prerequisite for Broad-Spectrum CD8+ T Cell Immunity
Background: Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has developed variants escaping neutralization antibody immunity established against the original virus. An understanding of broad-spectrum adaptive immunity, including CD8+ T cell immunity to wide range of epitopes, could help translational efforts to improve coronavirus disease 2019 (COVID-19) prevention and therapy. However, there have been few direct studies in which such immunity exists in a population. Methods: We selected SARS-CoV-2-conserved structural peptides that are not prone to mutation as antigens for broad-spectrum CD8+ T cell immunity. Peripheral blood mononuclear cells (PBMCs) from unexposed healthy donors were stimulated with these peptides in vitro and CD8+ T cell-specific response was monitored. The conserved peptide-specific CD8+ T cells were sorted for T cell receptor (TCR) repertoire sequencing. The presence of specific complementary determining region 3 (CDR3) clones was analyzed in a healthy cohort. Results: For each structural protein, including S, E, M, N, the conserved peptides could potentially provide the largest number of major histocompatibility complex-I (MHC-I) epitopes in the Oriental and Caucasian populations. For conserved peptides from spike (S), envelope (E), membrane (M), nucleocapsid (N) proteins, we found that there were no cross-reactive memory T cells in the unexposed individuals. Instead, their T cells contain naïve TCR repertoire recognizing these conserved peptides. Using TCR sequencing and CDR3 clustering for the conserved peptides specific T cells, we found that the recovered patients had a higher proportion of TCR repertoire similar with that of specific CD8+ T cells in unexposed individuals. Meanwhile, CDR3 clones of the above T cells were widely present in the healthy population. Conclusions: This study provides evidence of broad-spectrum SARS-CoV-2 specific CD8+ TCR repertoire in unexposed healthy population, which is implicated in the development and implementation of broad-spectrum vaccines against COVID-19.

A Neoantigen-Based Peptide Vaccine for Patients With Advanced Pancreatic Cancer Refractory to Standard Treatment
[Background] Neoantigens are critical targets to elicit robust antitumor T-cell responses. Personalized cancer vaccines developed based on neoantigens have shown promising results by prolonging cancer patients’ overall survival (OS) for several cancer types. However, the safety and efficacy of these vaccine modalities remains unclear in pancreatic cancer patients. [Methods] This retrospective study enrolled 7 advanced pancreatic cancer patients. Up to 20 neoantigen peptides per patient identified by our in-house pipeline iNeo-Suite were selected, manufactured and administered to these patients with low tumor mutation burden (TMB) (less than 10 mutations/Mb). Each patient received multiple doses of vaccine depending on the progression of the disease. Peripheral blood samples of each patient were collected pre- and post-vaccination for the analysis of the immunogenicity of iNeo-Vac-P01 through ELISpot assay and flow cytometry. [Results] No severe vaccine-related adverse effects were witnessed in patients enrolled in this study. The mean OS, OS associated with vaccine treatment and progression free survival (PFS) were reported to be 24.1, 8.3 and 3.1 months, respectively. Higher peripheral IFN-γ titer and CD4+ or CD8+ effector memory T cells count post vaccination were found in patients with relatively long overall survival. Remarkably, for patient P01 who had a 21-month OS associated with vaccine treatment, the abundance of antigen-specific TCR clone drastically increased from 0% to nearly 100%, indicating the potential of iNeo-Vac-P01 in inducing the activation of a specific subset of T cells to kill cancer cells. [Conclusions] Neoantigen identification and selection were successfully applied to advanced pancreatic cancer patients with low TMB. As one of the earliest studies that addressed an issue in treating pancreatic cancer with personalized vaccines, it has been demonstrated that iNeo-Vac-P01, a personalized neoantigen-based peptide vaccine, could improve the currently limited clinical efficacy of pancreatic cancer.

Deep Sequencing of the T-cell Receptors for Monitoring Peripheral CD8+ T Cells in Chinese Advanced Non-small-cell Lung Cancer Patients Treated with anti-PD-L1 antibody
Background: Atezolizumab, a high-affinity engineered human anti-PD-L1 antibody, has produced a clinical benefit for patients with advanced non-small-cell lung cancer (NSCLC). However, associated with T cell regulation, the immuno-modulatory effect of PD-L1 blockade and its biomarker in peripheral immunity remains illustrated. Methods: In a prospective cohort with 12 Chinese advanced NSCLC patients who received atezolizumab 1200 mg every three weeks as a second-line treatment, blood samples were obtained before and six weeks after atezolizumab initiation, and when disease progression confirmed. Patients were classified into a response or progression group according to Response Evaluation Criteria in Solid Tumors (RECIST)1.1. Fresh peripheral blood mononuclear cells (PBMCs) from patients were stained with anti-human CD3, CD8, and PD-1 antibodies for flow cytometry analysis. T cell receptor (TCR) β chains of CD8+ T cells were analyzed by Next-Generation Sequencing (NGS) at the deep level. Diversity, clonality, and similarity of TCR have been calculated before and after treatment in both groups. Results: Clonal expansion with high PD1 expression was detected in all patients’ peripheral CD8+ T cells before the treatment of atezolizumab. Unlike the progression group, the diversity of TCR repertoire and singletons in the TCRβ pool, increased over time with atezolizumab administration, and the TCR repertoire dynamically changes in the response group. The percentage of CD8+ PD1high terminal exhausted T cells declined in the response group after the PD-L1 blockade. Two patterns of TCR changes among patients who receive PD-L1 targeted immunotherapy were observed. Conclusions: Deep sequencing of the T-cell receptors confirmed the existence of CD8+ PD1high T cells with an exhaustion phenotype in Chinese NSCLC patients. Our study demonstrated that efficient anti-PD-L1 therapy could reshape the TCR repertoire for anti-tumor. Furthermore, singleton frequency may help us select patients who are sensitive to anti-PD-L1 immunotherapy.

Profiling T cell receptor β-chain in responders after immunization with recombinant hepatitis B vaccine
Background: T cells with edited T cell receptor β-chain variable (TRBV) involve in the immune response to recombinant hepatitis B surface antigen (rHBsAg) vaccine and the production of hepatitis B surface antibody (HBsAb). The immune repertoire (IR) profile and mechanism of vaccination positive responder (VPR) with rHBsAg is not fully understood. Methods: The IR of 6 VPRs (HBsAb+, HBsAg-) with rHBsAg vaccination was established by high throughput sequencing (HTS) technique and bioinformatics analysis, and comparing to those in 5 vaccination negative responders (VNRs) (HBsAb-, HBsAg-) who were also inoculated with rHBsAg. The repertoire features of BV, BJ, and V (CDR3)J genes selected and immune diversity in peripheral blood mononuclear cells (PBMC) of each subject were analyzed, respectively. Results: There is no significant difference in sequencing amplification indexes of each sample. However, there are TRBV15, BJ2-3 with significantly high expression levels in VPR compared with those in VNR group (both P <0.05). Further results show that BV15/BJ2-5 level was significantly increased from VPR than that of VNR group. Interestingly, the motif of CDR3 in TRBV15/BJ2-5 is mostly expressed as "GGETQ" or "GETQ". Additionally, there is no remarkable difference between the two groups of distribution of different clone expressed levels of V (CDR3) J. Conclusions: The features of IR in the VPR and VNR will contribute to explore the mechanism of positive response to rHBsAg, and contribute to development of optimized hepatitis B vaccine, and also provide partial interpretation of the VNR who is relatively few infected with HBV. Keywords: Complementarity determining region 3; Hepatitis B vaccine; High throughput sequencing; TR repertoire; Vaccination positive responder.
Rapid isolation and immune profiling of SARS-CoV-2 specific memory B cell in convalescent COVID-19 patients via LIBRA-seq
B cell response plays a critical role against SARS-CoV-2 infection. However, little is known about the diversity and frequency of the paired SARS-CoV-2 antigen-specific BCR repertoire after SARS-CoV-2 infection. Here, we performed single-cell RNA sequencing and VDJ sequencing using the memory and plasma B cells isolated from five convalescent COVID-19 patients, and analyzed the spectrum and transcriptional heterogeneity of antibody immune responses. Via linking BCR to antigen specificity through sequencing (LIBRA-seq), we identified a distinct activated memory B cell subgroup (CD11chigh CD95high) had a higher proportion of SARS-CoV-2 antigen-labeled cells compared with memory B cells. Our results revealed the diversity of paired BCR repertoire and the non-stochastic pairing of SARS-CoV-2 antigen-specific immunoglobulin heavy and light chains after SARS-CoV-2 infection. The public antibody clonotypes were shared by distinct convalescent individuals. Moreover, several antibodies isolated by LIBRA-seq showed high binding affinity against SARS-CoV-2 receptor-binding domain (RBD) or nucleoprotein (NP) via ELISA assay. Two RBD-reactive antibodies C14646P3S and C2767P3S isolated by LIBRA-seq exhibited high neutralizing activities against both pseudotyped and authentic SARS-CoV-2 viruses in vitro. Our study provides fundamental insights into B cell response following SARS-CoV-2 infection at the single-cell level.

Natural evolution mechanisms of the IDH mutant glioma revealed by multi-omics sequencing of a long-term survivor
Results: The center and edge of the tumor were diagnosed as LGG and GBM, respectively. They shared the same trunk mutations including IDH, TP53, and ATRX. They both have mixing cell origins and they have a highly correlated methylation level at the probe level. CIC, BRCA2, and RPA4 mutation which occurred only in G4 with mutant allele frequency(MAF) higher than 15% may contribute to evolution. NAF1 of which the MAF increased by 70% and the mutant RNA reads nearly doubled in G4 may also involve in the evolution. In the pathway level, the MSP-RON pathway was strongly up-regulated in G4. Concomitant with the tumor evolution, we discovered enhanced inflammatory signals represented by the up-regulation of the NF-κB pathway and the recruitment of mast cell, of which the absolute cell proportion increased from 3.9% to 6.9%. Contradictorily, the adaptive immune response was suppressed as we found pathways associated with IL17, dendritic cells, and cytotoxic T cells were down-regulated and the infiltration level of CD4+ and CD8+ T cells were both decreased. As for the TCR profile, only about 2% of blood clonotypes were detected in the tumor microenvironment. Nevertheless, two clonotypes were found significantly expanded. Clone frequencies of them were 38% and 19% in G2, respectively. And these were 24% and 11% in G4, respectively. Conclusion: Mutation of CIC, BRCA2, RPA4, and NAF1 and activation of the MSP-RON pathway could promote glioma evolution under natural conditions. Increased inflammatory response and decreased adaptive immune response may also contribute to the process. Besides, two highly expanded TCR clonotypes discovered in this case may serve as a potential adoptive cell therapy source in glioma.

T cells expanded from PD-1+ peripheral blood lymphocytes share more clones with paired tumor-infiltrating lymphocytes
Both tumor-infiltrating lymphocytes (TIL) and PD-1+ peripheral blood lymphocytes (PBL) are enriched for tumor-reactive clones recognizing known and unknown tumor antigens. However, the relationship between the T-cell receptor (TCR)-β repertoires of the TIL and T cells expanded from paired PD-1+ PBL and whether T cells expanded from PD-1+ PBL can be used to treat patients with cancer as TIL substitutes remains unclear. Here we established a highly efficient protocol to prepare polyclonal T cells from PD-1+ PBL. A functional T-cell assay and tetramer staining revealed that cells from PD-1+ PBL were relatively enriched for tumor-reactive T cells. Furthermore, deep TCR-β sequencing data revealed that an average of 11.29% (1.32%-29.06%, p=0.015, n=8) tumor-resident clonotypes were found in T cells expanded from paired PD-1+ PBL, and the mean accumulated frequency of TIL clones found in T cells expanded from PD-1+ PBL was 35.11% (7.23%-78.02%, p=0.017, n=8). Moreover, treatment of four patients, who failed multi-line therapy and developed acquired resistance to anti-PD-1, with autologous T cells expanded from PD-1+ PBL combined with anti-PD-1 antibody elicited objective responses from three of them. These results indicate that T cells expanded from PD-1+ PBL share more clones with paired TIL and could be used to treat patients with cancer as TIL substitutes.

Restricted TcR β chain CDR3 clonotype is associated with resolved acute hepatitis B subjects
Background: T cells play an important role in the prognosis of hepatitis B virus (HBV) infection, and are involved in the seroconversion of a patient from HBsAb negative to positive. To compare the T-cell receptor β-chain variable region (TcRBV) complementarity-determining region 3 (CDR3) in subjects with or without hepatitis B surface antigen (HBsAg) convert to hepatitis B surface antibody (HBsAb), the TcRBV was determined using high throughput sequencing (HTS). Methods: The clonotype and diversity of CDR3 in peripheral blood mononuclear cells of subjects with resolved acute hepatitis B (AHB, HBsAb+, HBsAg-) (n = 5), chronic hepatitis B (CHB, HBsAb-, HBsAg+) (n = 5), and healthy controls (HC, HBsAb-, HBsAg-) (n = 3) were determined and analyzed using HTS (MiSeq). Results: The overlapping rate of CDR3 clones of any two samples in AHB group was 2.00% (1.74% ~ 2.30%), CHB group was 1.77% (1.43% ~ 2.61%), and HC group was 1.82% (1.62% ~ 2.12%), and there was no significant difference among the three groups by Kruskal-Wallis H test. However, among the top 10 cumulative frequencies of clonotypes, only the frequency of clonotype (TcRBV20–1/BD1/BJ1–2) in AHB group was lower than that of HC group (P < 0.001). Moreover, exclude the 10 top clonotypes, there are 57 markedly different frequency of clones between AHB and CHB groups (18 clones up, 39 clones down), 179 (180–1) different clones between AHB and HC groups, and 134 different clones between CHB and HC groups. With regard to BV and BJ genotypes, there was no significant different frequency among the groups. Furthermore, there was no significant difference in the diversity of TcRBV CDR3 among the three groups (P > 0.05). Conclusions: Thus, there are 57 TcRBV clonotypes that may be related to HBsAg seroconversion of AHB subjects, but the diversity of TcRBV CDR3 is not significantly related to the HBsAb positive status.

Efficacy and safety of sintilimab in combination with chemotherapy in previously untreated advanced or metastatic nonsquamous or squamous NSCLC: two cohorts of an open-label, phase 1b study
Combining chemotherapy with immunotherapy improves the therapeutic outcome for first-line (1L) patients with advance nonsmall-cell lung cancer (NSCLC). Two cohorts of a phase 1b study (NCT02937116) aimed to evaluate the safety and efficacy of sintilimab, a PD-1 inhibitor, plus chemotherapy in 1L patients with nonsquamous and squamous NSCLC (nsqNSCLC/sqNSCLC); and to identify potential biomarkers for treatment response. Treatment-naïve patients with nsqNSCLC were enrolled and intravenously given sintilimab (200 mg), pemetrexed (500 mg/m2), and cisplatin (75 mg/m2), every 3 weeks (Q3W) for 4 cycles in cohort D. Treatment-naïve patients with sqNSCLC were enrolled and intravenously given sintilimab (200 mg), gemcitabine (1250 mg/m2), and cisplatin (75 mg/m2), Q3W, for 6 cycles in cohort E. The primary objective was to evaluate the safety and efficacy of the treatment. The additional objective was to explore biomarkers for the treatment efficacy. Twenty-one patients with nsqNSCLC, and 20 patients with sqNSCLC were enrolled in cohort D and cohort E, respectively. By the data cutoff (April 17, 2019), 8 (38.1%) patients in cohort D and 17 (85.0%) patients in cohort E experienced grade 3–4 adverse events. The median follow-up duration was 16.4 months (14.8–23.0) in cohort D and 15.9 months (11.7–17.7) in cohort E. The objective response rate was 68.4% (95% CI 43.4%, 87.4%) in cohort D and 64.7% (95% CI 38.3%, 85.8%) in cohort E. Neither PD-L1 expression nor tumor mutation burden value was significantly associated with an improved treatment response. Sintilimab plus chemotherapy exhibited manageable toxicity and an encouraging antitumor activity in patients with nsqNSCLC and sqNSCLC.

Characterization of Variable Region Genes and Discovery of Key Recognition Sites in the Complementarity Determining Regions of the Anti-Thiacloprid Monoclonal Antibody
Sequence-defined recombinant antibodies (rAbs) have emerged as alternatives to hybridoma-secreted monoclonal antibodies (mAbs) for performing immunoassays. However, the polyploidy nature of hybridomas often leads to the coexistence of aberrant or non-specific functional variable region (VR) gene transcripts, which complicates the identification of correct VR sequences. Herein, we introduced the use of LC-MS/MS combined with next-generation sequencing to characterize VR sequences in an anti-thiacloprid mAb, which was produced by a hybridoma with genetic antibody diversity. The certainty of VR sequences was verified by the functional analysis based on the recombinant antibody (rAb) expressed by HEK293 mammalian cells. The performance of the rAb was similar to that of the parental mAb, with IC50 values of 0.73 and 0.46 μg/L as measured by ELISAs. Moreover, molecular docking analysis revealed that Ser52 (H-CDR2), Trp98, and Trp93 (L-CDR3) residues in the complementarity determining regions (CDRs) of the identified VR sequences predominantly contributed to thiacloprid-specific recognition through hydrogen bonds and the CH–π interaction. Through single-site-directed alanine mutagenesis, we found that Trp98 and Trp93 (L-CDR3) showed high affinity to thiacloprid, while Ser52 (H-CDR2) had an auxiliary effect on the specific binding. This study presents an efficient and reliable way to determine the key recognition sites of hapten-specific mAbs, facilitating the improvement of antibody properties.

A pan-cancer clinical study of personalized neoantigen vaccine monotherapy in treating patients with various types of advanced solid tumors
Purpose: Due to their high tumor specificity and immunogenicity, neoantigens have been considered as ultimate targets for cancer immunotherapy. Neoantigen-based vaccines have demonstrated promising efficacy for several cancer types. To further investigate the anti-tumor potentials for other types of solid tumors, we designed a peptide-based neoantigen vaccine, iNeo-Vac-P01, and conducted a single-arm, open-labeled, investigator-initiated clinical trial (NCT03662815). Experimental Design: Personalized neoantigen vaccines were designed and manufactured according to our bioinformatics analysis results from the whole-exome sequencing (WES) of tumor and peripheral blood cell DNAs. Patients were scheduled to be vaccinated subcutaneously (s.c.) with adjuvant on days 1, 4, 8, 15, 22 (prime phase), and days 78, 162 (boost phase). Additional immunizations were administrated every 2~3 months as per patient's potential benefit. The safety and efficacy were assessed through adverse events, progression-free survival (PFS), overall survival (OS) and other parameters. Results: Of the 22 patients enrolled with advanced malignancies, twenty had no or mild adverse events, while two had grade 3 or 4 acute allergic reactions only after their 6th boost vaccination. The disease control rate (DCR) was 71.4%. The median PFS was 4.6 months, whereas the median OS was not reached (12-month OS=55.1%). Around 80% of individual peptides or peptide pools elicited measurable specific immune response. Additionally, our findings revealed several potential biomarkers for the prediction of better response. Conclusions: iNeo-Vac-P01 as monotherapy is feasible and safe for patients with advanced solid tumors. It could elicit T cell-mediated immune response targeting tumor neoantigens, and might have promising antitumor efficacy.
Restricted TcR β chain CDR3 Clonotype Is Associated with Resolved Acute Hepatitis B (AHB) patients with HBsAb
Background: T cells play an important role in the prognosis of HBV infection, and are involved in the seroconversion of a patient from HBsAb negative to HBsAb positive. To compare the T-cell receptor beta chain variable region (TcRBV) complementarity-determining region 3 (CDR3) in subjects with or without HBsAg seroconversion to HBsAb, the TcRBV was determined using high throughput sequencing (HTS). Methods: Clonotype and diversity of CDR3 in peripheral blood mononuclear cells (PBMCs) of subjects with resolved acute hepatitis B (AHB, HBsAb+, and HBsAg-) (n = 5), chronic hepatitis B (CHB, HBsAb-, and HBsAg+) (n = 5), and healthy controls (HC, HBsAb-, and HBsAg-) (n = 3) were determined and analyzed using HTS. Findings: The overlapping rate of CDR3 clones for any two samples in the AHB, CHB, and HC groups were 2.00% (1.74–2.30%), 1.77% (1.43–2.61%), and 1.82% (1.62%–2.12%), respectively; no significant difference was found among the three groups by KruskalWallis H test. In addition, among the top 10 cumulative frequencies of clonotypes, only the frequency of clonotype (TcRBV20-1/TcRBD1/TcRBJ1-2) in the AHB group was lower than that in the HC group (P < 0.001). Additionally, excluding the 10 top clonotypes, there were 57 markedly different frequencies of clones between the AHB and CHB groups. Furthermore, there was no significant difference in the diversity of TcRBV CDR3 among the three groups (P<0.05). InterpretationL Thus, there are 57 TcRBV clonotypes that may be related to HBsAg seroconversion in AHB subjects, but the diversity of TcRBV CDR3 is not significantly related to the HBsAb positive status.

Quantitative Analysis of Thymus-Independent Donor-Derived T Cell Expansion in Transplant Patients
Although thymus-independent donor-derived T cell expansion may determine the occurrence of graft-versus-host disease (GVHD) and relapse after transplantation, the characteristics and dynamics of the expansion process remain unclear. To address this issue, we monitored T cell receptor β repertoire at day 0, day 28, and day 61 after transplantation in 30 patients with hematologic malignancies by next-generation sequencing. The clonality index showed an increasing clonality over time (P = .001). The top 200 clonotypes accounted for more than half of the total clonotypes (median frequency, 63.55%) at day 61, and there was a remarkable overlapping between the top 200 clonotypes of each repertoire and its former repertoire (>50%). A normalized index, called the T Cell Response Index (TCRI), was designed on the basis of rank-shift analysis to quantify antigen-driven expansion. The TCRI during the first month was not related to relapse or GVHD (P> .05), whereas the TCRI during the second month was related to relapse (P = .006). Recipients with a TCRI below 1.0 during the second month had a higher cumulative relapse rate (31.25% versus 0%, P = .0323) and had a lower 1-year survival rate (56.25% versus 78.57%, P = .281). The clonotypes with strong competitiveness in the second month in the nonrelapse group preferentially used TRBV2, TRBV12-3, TRBJ1-1 and TRBJ1-5 segments (P< .01). In conclusion, homeostatic expansion predominates in the first month due to nonspecific T cell proliferation, whereas antigen-driven expansion predominates in the second month and results in a graft-versus-tumor (GvT) effect. Moreover, TCRI could serve as a quantitative indicator of GvT against relapse within the first year. The difference in V and J segment usage reveals that T cells responsible for potent GvT effect are similar among patients.

Engineering Magnetosomes for High-Performance Cancer Vaccination
A novel cancer vaccine is developed by using Fe3O4 magnetic nanoclusters (MNCs) as the core and cancer cell membranes decorated with anti-CD205 as the cloak. Because of the superparamagnetism and magnetization of MNCs, it is first achieved for the magnetic retention of vaccine in the lymph nodes with a magnetic resonance imaging (MRI) guide, which opened the time window for antigen uptake by dendritic cells (DCs). Meanwhile, the camouflaged cancer cell membranes serve as a reservoir of various antigens, enabling subsequent multiantigenic response. Additionally, the decorated anti-CD205 direct more vaccine into CD8+ DCs, facilitating the major histocompatibility complex (MHC) I cross-presentation. These unique advantages together lead to a great proliferation of T cells with superior clonal diversity and cytotoxic activity. As a result, potent prophylactic and therapeutic effects with few abnormalities are observed on five different tumor models. Therefore, such a cancer-derived magnetosome with the integration of various recent nanotechnologies successfully demonstrates its promise for safe and high-performance cancer vaccination.

Quantitative characterization of T-cell repertoire alteration in Chinese patients with B-cell acute lymphocyte leukemia after CAR-T therapy
Chimeric antigen receptor (CAR) T-cell therapy has displayed potent anti-leukemia activity in acute lymphocytic leukemia (ALL), acting as a new ray of hope to refractory/relapsed patients. However, the influence of CAR-T therapy on host immune system has not been well elucidated. Thus, We applied high-throughput T cell receptor β chain sequencing to track the dynamic change of T-cell repertoire induced by CAR-T therapy in B-cell ALL patients. Six Chinese patients achieving complete remission were under observation, whose blood samples, bone marrow samples and infused CAR-T samples were collected at serial time points before and after CAR-T therapy. We observed decreased TCR diversity and increased clonality of T-cell repertoire in both peripheral blood and bone marrow after CAR-T administration. The persistent T cell clones in blood and bone marrow expanded following leukemic cell destruction and were barely detected in CAR T-cell pool. For the first time, our results demonstrated CAR-T therapy could stimulate the clonal proliferation of CAR-negative T cells in patients. Considering other groups’ animal results indicating that CAR-T therapy could facilitate the proliferation of tumor antigen-specific T cells and that the emergence of these T cell clones followed the destruction of leukemic cells, they are most likely tumor antigen-specific.

Deep sequencing of the t-cell receptors for monitoring peripheral CD8+T cells in advanced NSCLC Chinese patients treated with anti-PD-L1 antibody
Background: Atezolizumab, a high-affinity engineered human anti-PD-L1 monoclonal immunoglobulin-G1 antibody, has dramatically produced clinical benefit for patients with advanced non-small cell lung cancer (NSCLC). However, the biomarker and immunomodulatory effects of PD-L1 blockade in peripheral immunity remains to be illustrated. Methods: In a prospective cohort with twelve Chinese advanced NSCLC patients who received intravenous atezolizumab 1200 mg once every 3 weeks as second-line treatment. Blood samples were obtained prior to and 6 weeks after atezolizumab initiation, and the time when disease progression confirmed. Patients were classified into response or progression group according to their clinical efficacy to atezolizumab. TCRβ chains of CD8+ T cells were analyzed by NGS using TCR profiling system at the deep level (ImmuQuad Biotech, Hangzhou China). Briefly, a 5' RACE unbiased amplification protocol was applied. Sequencing was performed on an Illumina MiSeq system. Meanwhile, fresh PBMCs from patients were stained with anti-human CD3, CD8, and PD1 antibodies for flow cytometry analysis. Results: Clonal expansion with high PD1 expression was detected in all patient peripheral CD8+ T cells before atezolizumab therapy. Unlike progression group, diversity of TCR repertoire and singletons in TCRβ pool, apparently increased overtime with atezolizumab administration and the TCR repertoire showed dynamic changes by calculated with Baroni-Urbani & Buser similarity index in response group. Meanwhile, the frequency of CD8+PD1high terminal exhausted T cells declined in response group after treatment. Conclusions: Results from deep sequencing of the T-cell Receptors highly indicated the existing of tumor antigen-specific T cells with exhaustion phenotype caused by persistent tumor antigen stimulation in Chinese advanced NSCLC patients. Our study demonstrated that an efficient anti-PD-L1 therapy could reshape T-cell repertoire for targeting and adapting to tumor heterogeneity, and reinvigorate the progenitor exhausted T cell population which includes tumor antigen-specific T cells for fighting tumor cells.

Chimeric Antigen Receptor T-Cell Therapy Shapes T Cell Repertoire in Chinese Patients with B-Cell Acute Lymphocyte Leukemia
Introduction: Chimeric antigen receptor (CAR) T-cell therapy has displayed potent anti-leukemia activity in refractory/relapsed acute lymphocytic leukemia (ALL). However, the influence of CAR-T therapy on host systemic and local immunity has not been well examined. We therefore applied high-throughput T cell receptor β chain sequencing to track dynamic change of T-cell repertoire in vivo induced by CAR-T therapy in B-cell acute lymphocytic leukemia patients. Patients and Methods: Six patients with 45 samples were under observation. The samples obtained to be tested were the peripheral blood mononuclear cell (PBMC) samples and bone marrow mononuclear cell (BMMC) samples before and after CAR-T administration, as well as the CAR-transduced autologous T cell samples on the day when they were to be infused to patients. The information of samples and patients was summarized in Table 1. The TCR full length mRNAs of these samples were deeply sequenced using the ImmunHub® TCR profiling system (ImmunQuad Biotech). Briefly, a 5'RACE unbiased amplification protocol was used. An algorithm was applied to raw sequencing data for PCR and sequencing errors correction and V, D, J, C gene segments mapping with IMGT®. The inverse Simpson index and the clonality index was calculated to estimate TCRβ clonotype diversity and the state of clonal proliferation of T cells. The donut chart and clone tracking heat map were generated by R.
